Single-cell integration benchmarking
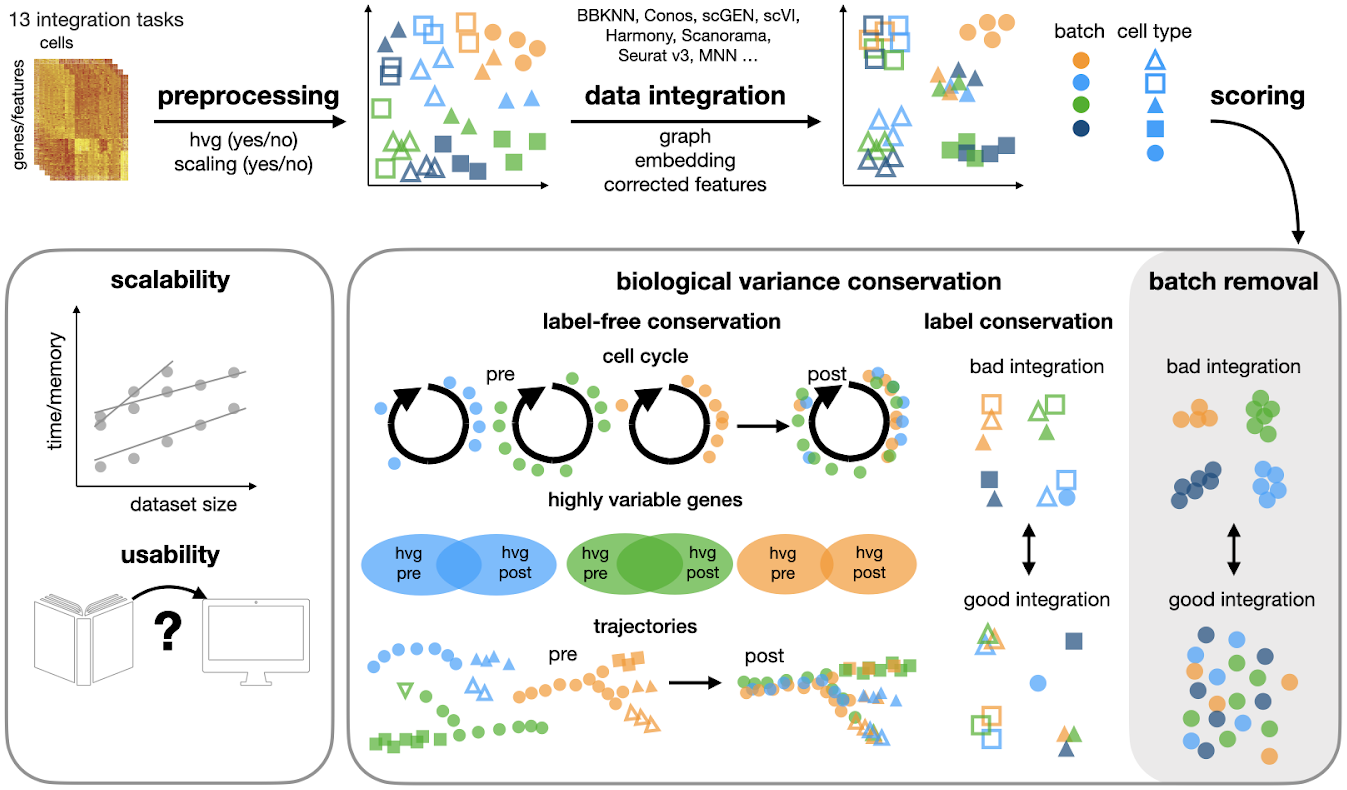
Single-cell integration benchmarking (scIB) is a project to assess the performance of scRNA-seq batch integration methods. We have used 14 metrics to evaluate 16 methods on 7 scRNA-seq (5 real and 2 simulated) and 6 scATAC-seq tasks. These metrics are designed to test both batch correction and conservation of biological variance.
For each task we also consider different combinations of pre-processing, including highly variable gene (HVG) selection and scaling. The usability and scalability of methods are also assessed.
Results
Here you can view all the individual benchmarking metrics broken down by dataset and method.
Datasets
Methods
ATAC Datasets
Usability
You can view the usability scores for each method here.
Links
More information about the project can be found at these links:
- scIB package - Python module implementing the metrics and wrapping the methods used in the evaluation.
- scIB pipeline - Snakemake pipeline implementing the evaluation workflow.
- scIB reproducibility - Additional code used to produce the results in the paper (including this website).
Citation
If any part of the project is useful for your work please cite:
Luecken MD, Buttner M, Chaichoompu K, Danese A, Interlandi M, Mueller MF, Strobl DC, Zappia L, Dugas M, Colome-Tatche M, Theis FJ. “Benchmarking atlas-level data integration in single-cell genomics” bioRxiv. 2020 DOI: 10.1101/2020.05.22.111161